Roche Cancer Research Application: High-resolution Melting Point Analysis for Detection of Methylation in Frozen Tissue Samples

1 Introduction
Promoter hypermethylation is a very common phenomenon in the transcription of malignant tumor genes and is considered as one of the hallmarks of malignant transformation of cells. DNA methylation analysis is a genetic testing tool for early cancer detection, risk assessment and therapeutic efficacy assessment with very attractive prospects.
The most common method for DNA methylation detection relies on the treatment of genomic DNA by sodium bisulfite, converting cytosine to uracil and unmodified 5-methylcytosine. The modified gene sequence is altered to identify methylated cytosine in subsequent PCR amplification products.
Through sulfite sequencing, the most accurate assessment of gene methylation can be performed to identify monomethylated cytosine. Several simpler PCR-based methods have been developed that are important for small-scale research laboratories. These methods are newer, simpler, and easier to operate. High resolution melting point analysis (HRM) is one of several methods. HRM is based on the "melting" nature of DNA in solution. The principle of the method is to distinguish disulfate-treated DNA templates with different methylated cytosine contents by melting point analysis based on different melting temperatures. HRM is a relatively simple and relatively inexpensive method because it does not require expensive probes and standardization of the reference gene. Through the application of HRM technology, all CpGs in the amplicon can be analyzed, and the homology and heterologous methylation can be distinguished according to the shape of the melting curve. This is very important because the methylation pattern of the promoter CpG island is usually non-homology.
Frozen tissue is a very important resource for biomarker detection. The performance of the main analytical methods was tested on formalin-fixed, paraffin-embedded (FFPE) tissue. The purpose of the current study was to evaluate and confirm the HRM analysis of promoter methylation in FFPE tissues from individuals with colorectal cancer.
2 Materials and methods
2.1 Control and study samples
CpGenome universal methylated DNA (Chemicon/Millipore, Billerica, MA) and DNA from peripheral blood mononuclear cells (PBMNCs) were used as methylation and non-methylation controls. Methylation control DNA was diluted in unmethylated PBMC DNA to generate methylation standards (50%, 25%, 10%, 5%, 1%, and 0.1%). After optimization of HRM testing, we analyzed frozen tissue samples from individuals with colorectal tumors.
2.2 DNA extraction and disulfate modification
PBMNC DNA and DNA from healthy volunteers from cultured tumor cell lines were isolated. For DNA isolates from cryopreserved tissue, sections of 10 μm thickness from FFPE tumor tissue blocks were used.
2.3 MethyLight detection
For a description of the MGMT and APC MethyLight tests, see the previous literature description. Briefly, PCR was performed on a LightCycler® 480 instrument.
2.4 high resolution melting point analysis
PCR amplification and HRM were performed on a LightCycler® 480 instrument (Roche) . Considering that FFPE DNA is highly fragmented, large amplicons cause a decrease in melting point resolution, and the amplicon is designed to not exceed 200 bp.
HRM data analysis was performed using Gene Scanning and the TM Calling software module (Roche). The melting curve is normalized by calculation of two normalized regions representing the pre-melting and post-melting of the PCR product. By this calculation, direct comparisons can be made to samples with different starting fluorescence intensities. The normalized temperature conversion melting curve shows that as the temperature increases, the fluorescence intensity decreases.
3 results and discussion
Standard dilutions were tested to determine the sensitivity of HRM detection. With all HRM assays (APC, MGMT, GSPT1 and PTEN), we were able to detect 1% methylated DNA in a non-methylated DNA background based on guaranteed reproducibility (Figure 1A-D) .
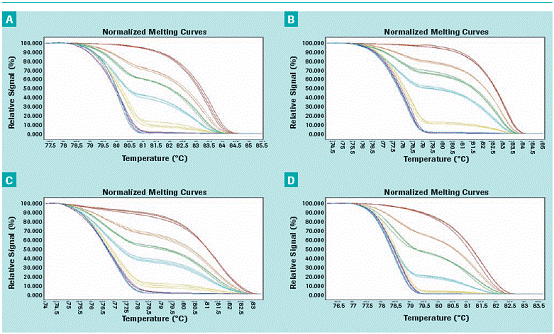
Figure 1 : Sensitivity of MGMT ( A ), APC ( B ), GSTP1 ( C ) and PTEN ( D ) methylation HRM detection.
The methylation standard curve is displayed in three ways. 100% methylation is shown in red, 50% methylation is shown in orange, 25% methylation is shown in green, 10% methylation is shown in cyan, 1% methylation is shown in yellow, 0.1% methyl The display is purple and 0% methylation is shown in blue.
Next, we evaluated the effect of formalin-fixed, wax-embedded treatment on promoter hypermethylation testing. We were able to detect the high methylation of the APC promoter with high detection reliability. Figure 2 (A–C) shows that the reproducibility and consistency of fresh and FFPE cell line DNA APC HRM assays are similar.
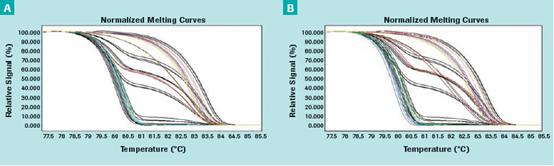
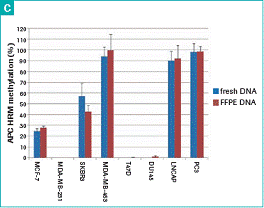
Figure 2: Using different tumor cell lines - methylation standards (black curve, 100%, 50%, 25%, 10%, 1%, 0.1%, 0%) and samples (color curves) Triple display. The methylation standard (black curve, 100%, 50%, 25%, 10%, 1%, 0.1%, 0%) and the sample (color curve) are shown in triple. (C) Inter-assay variability of fresh and FFPE tumor cell line DNA APC HRM assays from 8 breast and prostate cancer cell lines. The bar chart represents the mean and standard deviation of the six independent experiments.
The FFPE tissue promoter HRM methylation analysis has the following potential applications. The determination of methylation of primary tumors with lymph nodes and distant metastases not only elucidates epigenetic events that occur during disease progression, but also aids in the identification of predictive markers for paraffin-embedded tissue samples.
4 conclusion
Our study further confirmed the applicability of FFPE tissue promoter methylation analysis to quantify HRM. FFPE tissue is the largest source of sample material for normal controls and diseased tissues, and the application of these tissues is invaluable for research. Methodological evaluation is very important for the confirmation of experimental stability and can help to establish whether the test method is suitable as a research tool and possible routine detection means.
 Roche LightCycler ® 480 Real-Time PCR System "Three Good and One Fast"
A new experience seeks breakthroughs, innovative design interpretation of the classic!
    LightCycler® 480 is a classic automatic variable-flux real-time fluorescent PCR system that has been carefully developed and accumulated by Roche for more than ten years. The system includes temperature control module, xenon lamp source, optical detection system, supporting reagent supplies, and Dedicated data acquisition and analysis software. Its collection is flexible, fast and precise, providing an efficient research and diagnostic platform for gene expression and genetic analysis.

Good temperature control : The instrument's temperature control system introduces the exclusive patented Therma-base TM technology, which introduces a highly efficient liquid heat transfer layer between the heating and cooling elements with an unmatched inter-well temperature uniformity of ±0.1 °C. At present, the highest level of plate real-time quantitative PCR ensures good repeatability and uniformity of data, can achieve single-copy detection sensitivity, cover up to 11 orders of magnitude dynamic range, and can accurately distinguish as low as 1000 and The difference in concentration of 2000 copies, and the advanced application of HRM high-resolution melting curve analysis technology.
Good light source : Use wide-spectrum, high-intensity xenon lamp as the light source to achieve the best fluorescence excitation efficiency. With the unique five-edge reflection optical system, the optical path length can be effectively lengthened, which can significantly eliminate the edge effect common to similar instruments. A cold CCD is selected to ensure the highest detection sensitivity.
Good detection : Fully compatible with fluorescent dyes commonly used in real-time fluorescent PCR experiments by any combination of excitation and emission wavelengths, giving you the freedom to choose any analysis mode. The supporting software is easy to operate and user-friendly, allowing users to easily and time-savingly process high-throughput experimental data. The data analysis mode can be flexibly switched to support functions such as absolute quantification, melting curve, relative quantification, automatic genotyping and high resolution melting curve analysis. Roche's unique PCR Amplification Efficiency Correction ( E- Method ) algorithm guarantees experimental accuracy and is in line with internationally recognized MIQE standards.
Fast response : 384-well plates complete 40 cycles for less than 50 minutes, and 96-well plates complete 40 cycles for less than 30 minutes.
 Â
Roche offers a wide range of test kits - LightCycler® 480 SYBRIMaster, LightCycler® 480 Probe Master, LightCycler® 480 Genotyping Master,  LightCycler® 480 HRM Master, etc., is widely used in various fields such as nucleic acid quantification, gene expression, genotyping, methylation analysis, mutation site screening, chip detection and verification, and SNP locus discovery. Bureau of SFDA medical device registration certificate. To date, more than 25,000 related literature reports have been published.
LightCycler® 480 HRM Master, etc., is widely used in various fields such as nucleic acid quantification, gene expression, genotyping, methylation analysis, mutation site screening, chip detection and verification, and SNP locus discovery. Bureau of SFDA medical device registration certificate. To date, more than 25,000 related literature reports have been published.
 LightCycler® 480 HRM Master, etc., is widely used in various fields such as nucleic acid quantification, gene expression, genotyping, methylation analysis, mutation site screening, chip detection and verification, and SNP locus discovery. Bureau of SFDA medical device registration certificate. To date, more than 25,000 related literature reports have been published.
LightCycler® 480 HRM Master, etc., is widely used in various fields such as nucleic acid quantification, gene expression, genotyping, methylation analysis, mutation site screening, chip detection and verification, and SNP locus discovery. Bureau of SFDA medical device registration certificate. To date, more than 25,000 related literature reports have been published. Roche's strong marketing team, technical support and engineering team are located throughout the China, located in the Asia Pacific Technical Support Center at the Shanghai headquarters, through the free technical service hotline, the first time to respond to any questions you need to answer during use. Provide superior pre-sales product consultation, after-sales technical support, and accelerate your research process with the best solution. Intuitive and professional training course, hand-guided to quickly enter the role, become an expert in real-time PCR software and hardware operations.
Roche LightCycler® 480 Real-Time Fluorescence PCR System: Easy to operate, accurate, and reproducible, it's the smart choice for real-time PCR experiments.
Manual Recliner,Manual Recliner Chair,Best Manual Recliners,Best Manual Recliner
Jiangyin Kaili Health-Care Equipment CO., LTD. , https://www.kailimed.com